A walk through 2019

Scientific Highlights
1 Animal biodiversity and evolution
Conservation genomics to protect the Pyrenean desman.
IBE researchers led by José Castresana have developed a novel genomics methodology to analyse the networks of kinship relationships in the Pyrenean desman (Galemys pyrenaicus) across potential barriers to dispersal. The researchers have analysed the kinship of 70 specimens to identify the impact of different anthropogenic and natural river barriers on the dispersal of Pyrenean desman. The method could be of help in the development of conservation plans for aquatic and semi-aquatic endangered species.
Article: L. Escoda, A. Fernández-González, J.Castresana; 2019. Quantitative analysis of connectivity in populations of a semi‐aquatic mammal using kinship categories and network assortativity. Molecular Ecology Resources 19, 310-326. DOI: 10.1111/1755-0998.12967

Image of the Pyrenean desman. Credit: Jorge González-Esteban.
Paving the way towards cryptic species delimitation.
Mitochondrial DNA (mtDNA) sequencing has led to an unprecedented rise in the identification of cryptic species. However, it is widely acknowledged that nuclear DNA (nuDNA) sequence data are also necessary to properly define species boundaries. In this study, IBE researchers led by Roger Vila have focused on the intriguing case of the butterfly Thymelicus sylvestris (Lepidoptera: Hesperiidae), a small orange butterfly suspected to hide cryptic species. The researchers found a strong mtDNA/nuDNA discrepancy among populations, which could only be explained by long periods of geographic isolation followed by range expansions, homogenizing the nuclear but not the mitochondrial genome. The study concludes that species delimitation must be addressed by combining several and complementary types of molecular markers, as mtDNA and nuDNA do not necessarily follow the same mechanisms of evolution and their respective evolutionary history may reflect complementary aspects of past demographic and biogeographic events.
Article: JC. Hinojosa, D. Koubínová, MA. Szenteczki, C. Pitteloud, V. Dincă, N. Alvarez & R. Vila. (2019) A mirage of cryptic species: Genomics uncover striking mitonuclear discordance in the butterfly Thymelicus sylvestris. Molecular Ecology. DOI: 10.1111/mec.15153

The butterfly Thymelicus sylvestris stands out as a species with extreme genetic diversification in the mitochondrial DNA. Credit: Vlad Dinca.
2 Comparative and computational genomics
The genomic history of the Iberian Peninsula reconstructed.
An international team led by IBE researcher Carles Lalueza-Fox has developed a genetic map of the Iberian Peninsula covering the last 8,000 years. The findings show an invasion of steppe descendants who replaced almost all men 4,000 years ago. In the study, the team analysed the genomes of 271 inhabitants of the peninsula from different historical periods and has contrasted them with data collected in previous studies of 1,107 ancient and 2,862 modern subjects. The results show an unprecedented image of the transformation of the Iberian population along different historical and prehistoric stages.
Article: I. Olalde, […], C. Lalueza-Fox; The genomic history of the Iberian Peninsula over the past 8000 years. 2019. Science. DOI: 10.1126/science.aav1444

One of La Braña's brothers, with the skeleton mounted. They are without a doubt the oldest brothers genetically detected. Credit: Julio Manuel Vidal Encinas.
The oldest molecular information to date illuminates the history of the extinct Gigantopithecus.
A team led by IBE researcher Tomàs Marquès-Bonet and Enrico Cappellini from the Globe Institute has rebuilt for the first time proteins from approximately two million-year-old fossil remains. This is the first molecular evidence that allows clarifying the evolution of the human and great apes lineage beyond what the ancient DNA techniques allow. The method used, which has solved the evolutionary history of the extinct Gigantopithecus blacki from a molar fossil, could be used in the analysis of many other ancient remains, shedding light on the evolutionary history and origin of hominids.
Article: F. Welker, J. Ramos-Madrigal, M. Kuhlwilm, W. Liao, P. Gutenbrunner, M. de Manuel, D. Samodova, M. Mackie, M.E. Allentoft, A.-M. Bacon, J. Cox, C. Lalueza-Fox, J.V. Olsen, F. Demeter, W. Wang, T. Marques-Bonet, E. Cappellini; 2019; Dental enamel proteome shows that Gigantopithecus was an early divergent pongine. Nature 576: 262-265

Artistic representation of Gigantopithecus blacki. Credit: Kumi Kayama
3 Complex systems
Statistical physics of liquid brains.
Liquid neural networks (or ‘liquid brains’) are a widespread class of cognitive living networks characterized by a common feature: the agents (ants, termites or immune cells, for example) move in space. Thus, no fixed, long-term agent-agent connections are maintained, in contrast with standard neural systems. In this study, IBE researchers led by Ricard Solé have explored the collective dynamics, memory and learning properties of liquid brains under the perspective of statistical physics. The study shows that, despite their intrinsic physical differences, liquid brains share key properties with standard neural systems – like the use of fluctuations in ways similar to those found in cortical networks -, but strongly depart in other ways which may be crucial for their cognitive potential and ability to solve particular problems. Understanding the cognitive complexity of liquid brains and its limits can provide deep insights into the evolution of information-processing, computational systems grounded in living structures.
Article: J. Piñero, R. Solé. Statistical physics of liquid brains; 2019; Phil. Trans. R. Soc. B; DOI: 10.1098/rstb.2018.0376

4 Functional genomics and evolution
A crucial mechanism for multicellularity is highly conserved in closest unicellular relative of animals.
IBE Researchers led by Iñaki Ruiz-Trillo have identified that a unicellular relative of animals undergoes a transient stage resembling an epithelium tissue. The process, observed during coordinated cellularization of the protist Sphaeroforma arctica, resembles cellularization of coenocyte in animals, a specialized form of cytokinesis that results in the formation of a polarized epithelium during early embryonic development. Researchers have shown that the polarized layer of cells in Sphaeroforma arctica relies, as in animals, on an actomyosin network to hold the transient structure and that the temporary stage goes in hand with the activation of genes involved in cell adhesion.
Article: Dudin O, Ondracka A, Grau-Bové X, Haraldsen AAB, Toyoda A, Suga H, Bråte J, Ruiz-Trillo I. A unicellular relative of animals generates a layer of polarized cells by actomyosin-dependent cellularization. eLife 8:e49801. doi: 10.7554/eLife.49801.

Image of Sphaeroforma arctica just before cellularization, forming a structure reminiscent of an epithelium. In green, the actin network. In blue, the cell nuclei. Credit: Omaya Dudin.
The onset of metamorphosis depends on only one gene.
IBE researchers led by David Martín and Xavier Franch have identified the gene responsible for initiating insect metamorphosis. The gene involved, E93, was known to promote the formation of adult structures during metamorphosis, but it was unknown that it was the only one essential to initiate this process. The study confirms that the expression of E93 can initiate the metamorphosis of the insect even when it has not yet reached the final stages of its growth.
Article: S. Chafino, E. Ureña, J. Casanova, E. Casacuberta, X. Franch-Marro, D. Martín. Upregulation of E93 Gene Expression Acts as the Trigger for Metamorphosis Independently of the Threshold Size in the Beetle Tribolium castaneum; Cell Rep. 2019 Apr 23;27(4):1039-1049.e2. Doi: 10.1016/j.celrep.2019.03.094.

Pupae of the flour beetle Tribolium castaneum initiating the metamorphosis after activating the E93 gene, based on different initial sizes. Credit: David Martín.
5 Population genetics
Artificial intelligence applied to the genome identifies an unknown human ancestor.
By combining deep learning algorithms and statistical methods, an international team led by IBE researcher Jaume Bertranpetit has identified, in the genome of Asiatic individuals, the footprint of a new hominid who cross bred with its ancestors tens of thousands of years ago. Modern human DNA computational analysis suggests that the extinct species was a hybrid of Neanderthals and Denisovans and crossbred with Out of Africa modern humans in Asia. This finding would explain that the hybrid found last summer in the caves of Denisova–the offspring of a Neanderthal mother and a Denisovan father–, was not an isolated case, but rather was part of a more general introgression process. The study uses deep learning for the first time ever to account for human evolution, paving the way for the application of this technology in other questions in biology, genomics and evolution.
Article: Mayukh Mondal, Jaume Bertranpetit, Oscar Lao. Approximate Bayesian computation with Deep Learning supports a third archaic introgression in Asia and Oceania; Nature Communications 10, Article number: 246 (2019) DOI: 10.1038/s41467-018-08089-7
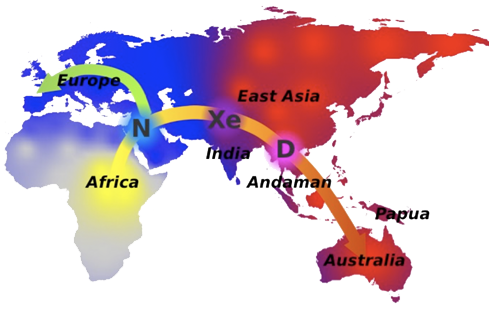
N: Neanderthal, D: Denisona Xe: new ancestral population, related to both Neanderthal and, mainly, Denisova. Credit: Oscar Lao
The genetic imprint of Palaeolithic populations has been detected in current North African populations.
A team led by IBE researcher David Comas has performed for the first time an analysis of the complete genome of the population of North Africa. In their analysis they have identified a small genetic imprint of the inhabitants of the region in Palaeolithic times in the genomes of present North Africans, thus ruling out the theory that recent migrations from other regions completely erased the genetic traces of ancient North Africans.
Article: G. Serra-Vidal, M. Lucas-Sanchez, K. Fadhlaoui-Zid, A. Bekada, P. Zalloua, D. Comas. Heterogeneity in Palaeolithic Population Continuity and Neolithic Expansion in North Africa, Current Biology (2019); DOI: 10.1016/j.cub.2019.09.050.
The year in numbers
 Staff
Staff
MEMBERS
STAFF

SCIENTIFIC STAFFby professional category
INTERNATIONAL
RESEARCHERS
ARE REPRESENTED
AT IBE
 Publications
Publications
RESEARCHERS
 Competitive and private funding
Competitive and private funding
INTERNATIONAL
RESEARCH PROJECTS