Highlighted papers
A 3-way hybrid approach to generate a new high quality chimpanzee reference genome (Pan_tro_3.0)
Kuderna, L.F.K.; Tomlinson, C.; Hillier, LD.W.; Tran, A.; Fiddes, I.; Armstrong, J.; Laayouni, H.; Gordon, D.; Huddleston, J.; Garcia Perez, R.; Povolotskaya, I.; Serres Armero, A.; Gómez Garrido, J.; Ho, D.; Ribeca, P.; Alioto, T.; Green, R.E.; Paten, B.; Navarro, A.; Bertranpetit, J.; Herrero, J.; Eichler, E.E.; Sharp, A.J.; Feuk, L.; Warren, L.C.; Marques-Bonet, T. (2017)
GigaScience 1;6(11):1-6 doi: 10.1093/gigascience/gix098
Genome reference sequences are an invaluable resource to study evolution, as they allow us to directly assess differences in life’s blueprint in individuals or species. Nevertheless, getting accurate and contiguous reference assemblies is far from trivial. Several new sequencing platforms and library preparation techniques for genome assembly have emerged in recent years, including long sequencing reads and chromosomal conformation maps. In this work, an array of these new techniques was benchmarked and integrated to generate a diverse set of orthogonal sequencing data of the chimpanzee genome, which allow us to generate the current version of the chimpanzee reference genome assembly, an essential tool to understand the evolution of our species. This way, over 164 Mbp of new repeat sequence were added, 77% of gaps present in the previous chimpanzee assembly were closed and 851 Kbp of additional exonic sequences were resolved.
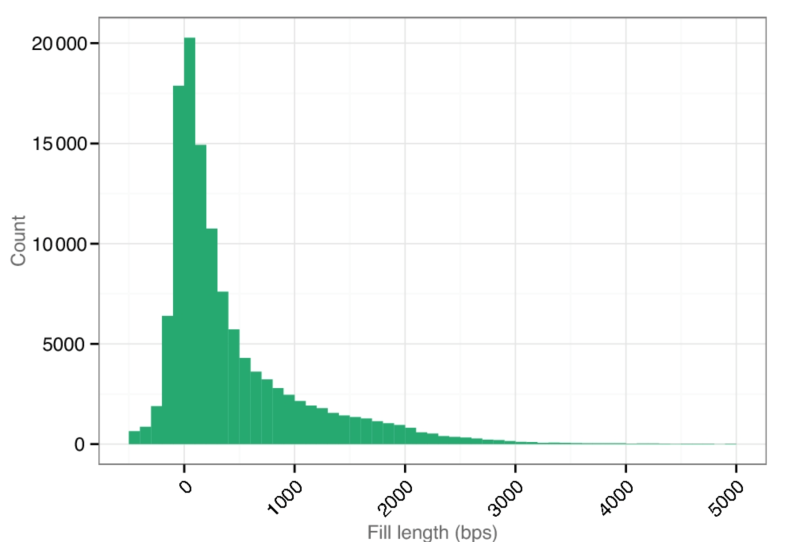
Length distribution of filled gaps (77%) compared to the previous chimpanzee reference assembly.
Go to the official publication
